|
Browse to the downloaded tutorial1-linksets directory to select the linksets used for extension.The MTI linksets support miRBase accession IDs, so we can select the miRBase ID column. The network attribute is the column that is used to identify the entities in the linkset and needs to contain one of the supported identifiers. The user network is the network you want to extend. Select the correct elements in the drop-down boxes for user network and network attribute.Go to ‘Apps -> CyTargetLinker -> Extend network’ (see Figure 4).Let’s start adding the known targets of the two microRNAs of interest.Install the CyTargetLinker app via the app manager (Apps -> App Manager -> Select CyTargetLinker -> Click ‘Install’).Step 3: Built regulatory network in Cytoscape 
Download the zip file and unzip it so both xgmml files are in the tutorial1-linksets directory.For this tutorial, you can download a zip file containing both linksets from here. In this example, we will use microRNA-gene interaction linksets from miRTarBase (validated interactions) and TargetScan (predicted interactions). A user can also easily create their own linksets.īefore starting CyTargetLinker you need to download the linksets of interest and store them in a directory. CyTargetLinker is not limited to the provided linksets. Several linksets for different use cases, species and interaction types are provided on our website, see Figure 3. Each linkset contains interactions consisting of two nodes, source and target, connected through one directed edge. The networks are stored in XGMML (the eXtensible Graph Markup and Modeling Language) format, which is supported by Cytoscape. This could be regulatory interactions like microRNA-gene interactions. The resulting biological network of two microRNAs from the example data, as shown in Figure 2.Ī linkset is a network containing additional information about and links between nodes in the network. After clicking ‘OK’, you will receive the following warning: “No edges will be created in the network the source column is not selected.Specify the microRNA column as the Source Node (click on header and select green circle) and the miRBase ID as a Source Node Attribute (green data sheet), as shown in Figure 1.Select the downloaded microRNA.xlsx file. Open the datafile containing the microRNAs in Cytoscape via File -> Import -> Network from File.The datafile should contain at least two columns, (i) the microRNA name and (ii) the corresponding miRBase accession number.Īn example datafile, in which two microRNAs known to be involved in obesity are listed, can be downloaded from here ( microRNA.xlsx). Step 1: Import a set of microRNAs into Cytoscapeįirst you will need to create a biological network in Cytoscape containing your microRNAs of interest. 
The tutorial was created for the following versions: 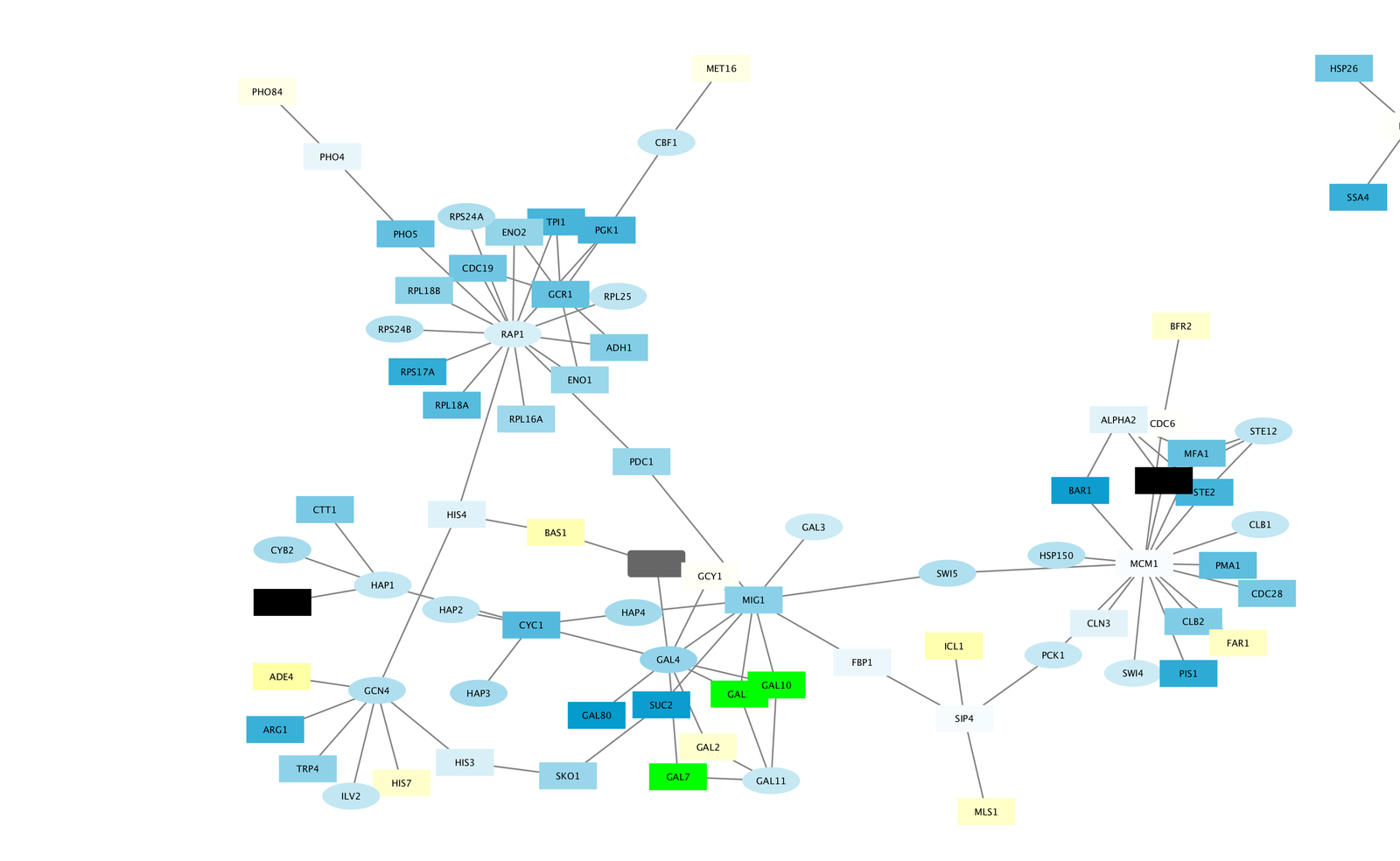
In this tutorial, a step-by-step explanation is given on how to extend a list of microRNAs with predicted and validated microRNA-target interactions (MTIs). The visualization options allow biological interpretation of complex regulatory networks in a graphical way. Tutorial 1 | CyTargetLinker - Extend your biological networks in Cytoscape Home Documentation Tutorials Link sets Contact Tutorial 1 Extend a set of microRNAs with target informationĬyTargetLinker provides a quick and extensive enrichment of networks in Cytoscape.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
